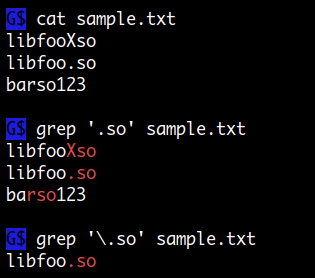
This command matches lines in the FASTA file that start with a ">" character, i.e. Grep -c '^>' mel-all-chromosome-r6.20.fasta One example of how regex can come in handy is using the ^ special character to quickly count how many sequences are in a FASTA file, which we would do as follows: Grep '\.' dmel-all-no-analysis-r6.20.gff will literally match a period. matching a period character "."), we would need to use a "\" to escape it first: Note that if we want to match any of these special characters literally (e.g. \: "escapes" meta-characters, allows literal matching : matches any characters *except* ones enclosed (note: is different from ^) : matches any character except new lines Regex has certain meta-characters that are reserved for special uses: Regex is extremely powerful, but can also get (very) complicated, so we'll just stick to a few basic uses. In other words, they allow you to match complex patterns with grep, not just exact matches. 
Regular expressions, aka "regex", are patterns that describe sets of strings. There are many other functions of grep! When in doubt, remember you can check the help page using man grep.or just by using google :) Pattern matching with regular expressions Grep -w -A 1 '>X' dmel-all-chromosome-r6.20.fasta We use the -w flag to match whole words, combined with -A to get the line after a match: Say we want to pull out a specific sequence from a FASTA file, like the X chromosome. Remember that we can combine different grep arguments with each other! E.g.: the command grep -f patterns.txt file.txt will match all patterns in patterns.txt against file.txt Grep -f pattern.txt matches a list of patterns contained in pattern.txt against target file. Note: grep -A is very useful for pulling out specific lines from a FASTA file.just make sure your FASTA file is single line and not multi-line! grep -C returns matching line and n lines before and after match grep -B returns matching line and n lines before match grep -A returns matching line and n lines after match Equivalent to grep 'pattern' file | wc -l Grep -c counts the number of lines that match the pattern. Grep -o returns only the matching words, not the entire line. Grep -v inverts, returning lines that do not match the pattern. In the above example, grep -i 'DOG' would still match the line Grep -w 'dog' sample.txt would match the string, but grep -w 'do' sample.txt would not. However, there are a huge number of arguments that can modify how grep behaves. The above three grep commands will all match the text in the example file and will print the line.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
